Duchenne Muscular Dystrophy (DMD) Model
Duchenne muscular dystrophy (DMD) is a genetic disorder characterized by progressive muscle degeneration and weakness due to the alterations of dystrophin.
Muscle tissue specification, development, and function in zebrafish are highly conserved with other vertebrates, including humans. In addition, the impact of disease and therapeutics can be studied much earlier due to its fast development. Transparent and accessible embryos allow for non-invasive tissue analysis, immunohistochemistry, and automated locomotion monitoring.
DMD model is designed for fast and high-throughput efficacy evaluation of candidate therapeutic compounds. This model allows identifying compounds that prevent muscle degeneration and the subsequent locomotor alterations.
Applications
- Study the causes and mechanisms of muscle impairments.
- High throughput screening of potential therapies for DMD.
Advantages
High conservation of zebrafish muscles with vertebrates and humans.
DMD muscle damage is already evident in embryos, which allows early studies of disease impact, and therefore, earlier drug efficacy testing.
Transparent embryos allow the optical birefringence-based evaluation of muscular integrity.
Automated locomotion monitoring highlights muscle functionality impairment rapidly.
Method description
Early-life ta222a dystrophin-null mutant embryos are exposed to the molecule of interest to determine mortality/teratogenicity and muscle integrity dose-effect curves. Muscle integrity is evaluated through optical birefringence, an optic phenomenon resulting from the diffraction of polarized light through the pseudo-crystalline array of the muscle sarcomeres. Zebrafish healthy muscles can be detected as a bright area in an otherwise dark background. Muscle damage, as seen in the ta222a mutant, can be detected by a reduction in the birefringence as shown in Figure 1. The phenotypic analysis must be coupled with genotypic homozygous confirmation to determine the percentage of homozygotes that recover the dystrophic phenotype after treatment with the compound of interest.
Readouts
- Muscle integrity: determined through optical birefringence.
- Genotyping: individual phenotype-genotype correlations in homozygous, heterozygous, and wt larvae.
- Locomotion activity (optional): traced and analyzed by DanioVision™ software (Noldus IT) automated video monitoring at 6 dpf.
 Figure 1. Analysis of muscle integrity through optical birefringence. A) Representative images of muscular phenotypes of wild-type, heterozygous and homozygous ta222a mutant zebrafish larvae at 6 dpf. B) Birefringence quantification by dark area calculation as a measure of muscle degeneration. The lack of dystrophin in ta222a mutants has an impact on muscle integrity compared to wild-type fish, only appreciated in homozygous mutants. Mean and SEM are plotted. ***p<0.001 vs. wild-type; ###p<0.001 vs. heterozygous.
Figure 1. Analysis of muscle integrity through optical birefringence. A) Representative images of muscular phenotypes of wild-type, heterozygous and homozygous ta222a mutant zebrafish larvae at 6 dpf. B) Birefringence quantification by dark area calculation as a measure of muscle degeneration. The lack of dystrophin in ta222a mutants has an impact on muscle integrity compared to wild-type fish, only appreciated in homozygous mutants. Mean and SEM are plotted. ***p<0.001 vs. wild-type; ###p<0.001 vs. heterozygous.
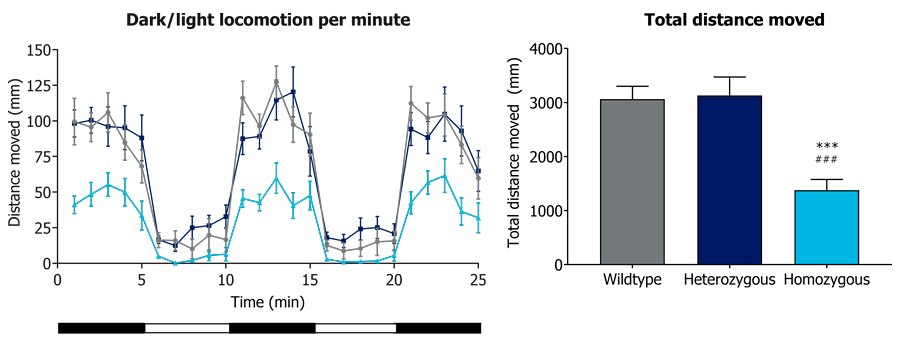 Figure 2. Locomotion activity assessment. A) Light/Dark cycles locomotion behavior per minute and B) total distance traveled by wild-type, heterozygous and homozygous ta222a mutant zebrafish larvae at 6 dpf. In both analyses, homozygous mutants move less compared to the other genotypes. Mean and SEM are plotted. **p<0.01 vs. wild-type; ###p<0.001 vs. heterozygous.
Figure 2. Locomotion activity assessment. A) Light/Dark cycles locomotion behavior per minute and B) total distance traveled by wild-type, heterozygous and homozygous ta222a mutant zebrafish larvae at 6 dpf. In both analyses, homozygous mutants move less compared to the other genotypes. Mean and SEM are plotted. **p<0.01 vs. wild-type; ###p<0.001 vs. heterozygous.
We'd like to hear from you
If you want more information about our DMD
models or have any other questions, please
contact our experts.
References
- Kawahara G, Kunkel LM. Zebrafish based small molecule screens for novel DMD drugs. Drug Discov Today Technol. 2013 Spring;10(1):e91-6.
- Jacoby AS, Busch-Nentwich E, Bryson-Richardson RJ, Hall TE, Berger J, Berger S, Sonntag C, Sachs C, Geisler R, Stemple DL, Currie PD. The zebrafish dystrophic mutant softy maintains muscle fibre viability despite basement membrane rupture and muscle detachment. Development. 2009 Oct;136(19):3367-76.